A growing number of bacterial strains are becoming resistant to common antibiotics. New types of antibiotics are needed, but the method we currently use to do that is outdated. This new method discovered by Leiden researchers can find DNA clusters in bacteria that can be used to make antibiotics.
New antibiotics
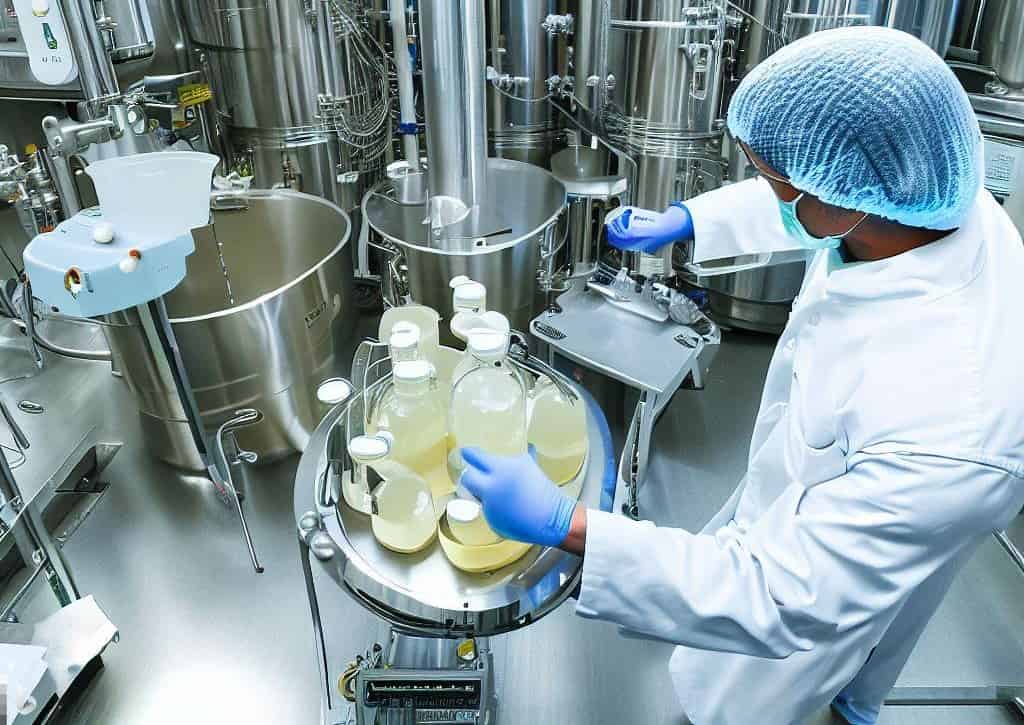
The quest for new antibiotics is becoming increasingly urgent. Smart software helps in this search by analyzing bacteria. The standard method of finding antibiotics is by using bacteria and fungi cultures that are thought to be capable of making antibiotics. Researchers grow microorganisms and see if the molecules can then inhibit bacteria. This smart software would replace this process.
Pristinine
Professor Gilles van Wezel from the Institute of Biology Leiden, set up the research together with Marnic Medema, University of Wageningen. The researchers developed the software that found 42 new types of clusters that met the stipulated conditions. These 42 clusters were able to later be developed into proteins with antibiotic activity. The researchers chose one of the clusters which best met the conditions, and made a small amount of pristinine, a potential new antibiotic, from it.
Pristinine is part of the lanthipeptides sub-class, which many antibiotics belong to. The usefulness of pristinine needs further research and the other 41 clusters are also undergoing further research.
The project was funded by the Topsector Chemie, a Dutch group that falls under the Netherlands Enterprise Agency. The project and the research results are seen as open source science. This means that the entire research is online and free for other research groups to take up.
Also interesting:
New, promising antibodies against SARS-CoV-2
Dutch TU Delft biotechnologists turn yeast into mini-factories for making medicine