Since the outbreak of the corona pandemic, research has been underway around the world to find cures and vaccines for the COVID-19 infection caused by the SARS CoV-2 virus. Hardly a day goes by without some laboratory reporting progress. Yet there are still no reliable solutions. Scientists at the Max Planck Institute for Biophysics in Frankfurt am Main (Germany) now want to analyze the surface protein of the novel coronavirus in order to find areas where vaccines and drugs can bind.
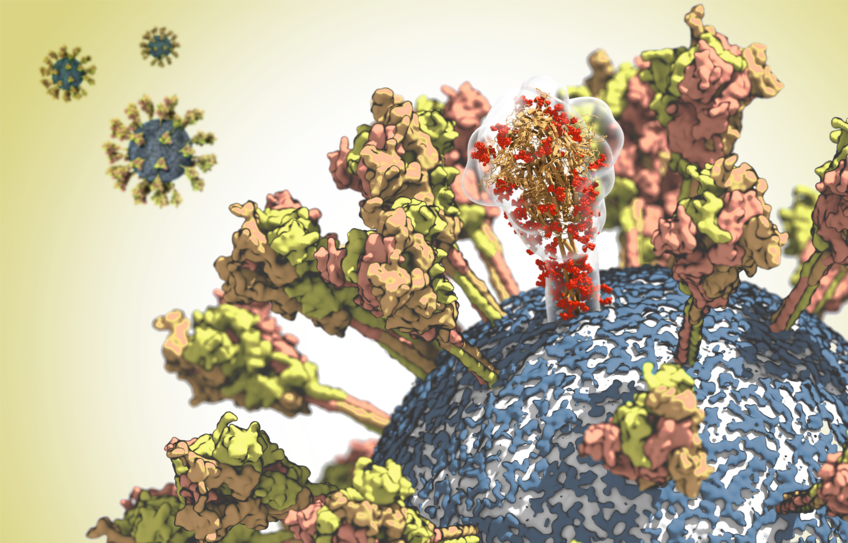
The corona virus – which quite literally means “crown virus” – owes its name to the glycoprotein spike. This molecule protrudes from the virus shell like the spikes of a crown. The researchers aim to study the structure of this protein in more detail as part of their analysis. And thereby identify potential targets suitable for antibodies and inhibitors. Precise information on where and how the SARS-CoV-2 virus could be attacked is an important prerequisite for the development of new vaccines and drugs.
Spike protein – weapon and Achilles’ heel at the same time
The spike protein is the weapon that the corona virus uses to attack cells. It binds primarily to a receptor called ACE2 located on human cell surfaces. This enables the virus to fuse with the cell membrane and release its genetic material into the cell interior. However, Max Planck scientists believe that it’s precisely this spike protein that could not only be the virus’s best weapon, but also its “Achilles’ heel.”
Due to its exposed position, i.e. how it protrudes from the virus shell, the spike protein also acts as a weak point for the immune system to attack. The researchers explain that antibodies could recognize the virus from its spike protein, bind to it and thereby mark it as a target for immune cells. However, the virus has a trick up its sleeve: it uses sugar molecules to try to hide its spike proteins from immune cells.
Sugar shield as a defense
To switch off this protective sugar shield, the scientists not only analyse the spike protein on its own, but also the virus’s membrane shell and protective shield. By doing so, they hope to be able to determine how the spike proteins move on the surface of the virus and how they change their shape – “with an accuracy that matches the size of an atom.”
Computations for this dynamic model of the spike protein are “extremely complex.” Nevertheless, they do reveal the most minute details of the protein structure. “To do this, we need the Max Planck Institute’s supercomputers with their enormous computational power,” explains Gerhard Hummer. He’s the director at the Max Planck Institute for Biophysics.
The purpose of the model is to detect domains where antibodies can reliably bind to as well as to discover places where inhibitors are able to bind. These will subsequently be compared with the binding properties of existing drugs with the help of a computer in order to identify drugs that can block the spike protein. “Of course, redesignating drugs that are already on the market is far quicker than finding new active substances and testing them in lengthy clinical trials,” said Hummer.
Cover photo: surface of the CoV-2 virus. One molecule of the spike protein is depicted translucently in order to highlight its complex spatial structure. © MPI f. Biophysics