The search for effective substances for medications and vaccines tends to be laborious and often in vain. Scientists looking for a vaccine to combat the SARS-CoV-2 coronavirus and medication to treat COVID-19 could now be helped by a new molecular library.
At the Helmholtz Zentrum Berlin for Materials and Energy (HZB), the MX team together with the Drug Design Group at the University of Marburg (both in Germany) have compiled a new molecular library. It is comprised of 1,103 organic molecules that could be used as building blocks for new drugs – not just to combat COVID-19 – and is now available to researchers worldwide.
Like a key in a lock
Medications normally have to dock to proteins in an organism before they can take effect. In the same way that a key must fit precisely into a lock, a medication requires part of the drug molecule to fit into depressions or cavities in the target protein. The team from the Department of Macromolecular Crystallography (MX) at HZB headed by Dr. Manfred Weiss. While he Drug Design Group is led by Prof. Dr. Gerhard Klebe of the University of Marburg.

These libraries are made up of tiny organic molecules (fragments), “whereby the functionally important cavities and depressions on the surface of proteins can be identified and mapped,” explain the scientists. “For this purpose, protein crystals are impregnated with the fragments and then analyzed with high-intensity X-ray light. This enables 3D structural details to be collected which have an atomic resolution. Among other things, this method can be used to determine how well a particular molecule fragment docks to the target protein.”
Successful first test
The draft of this type of ‘F2X Universal’ library of 1,103 substances has now been published by the MX team. “A representative selection of 96 compounds has been extracted from this library, known as the ‘F2X Entry Screen,'” the researchers state. “This selection has now been successfully tested by the HZB MX team at the MAX IV X-ray laboratory in Lund, Sweden, and at BESSY II (a synchrotron radiation source that produces extremely bright X-ray light housed at HZB in Germany) following this publication.
“The HZB and MAX IV teams have verified the efficacy of the F2X entry library in this study by screening the target enzymes endothiapepsin and the Aar2/Rnase-H complex.” As a next step, the researchers of the MX team intend to use the complete universal library.
“For this study, the fragment-screening experts at HZB-BESSY II in Berlin worked very closely with the FragMAX project team at MAX IV,” said Dr. Uwe Müller, from the MX team at HZB, who helped to establish both the three MX Beamlines at BESSY II and the BioMAX Beamline at MAX IV. “This enabled both partners to further develop their own technology platforms and use them for the imaging of the functional surfaces of a variety of proteins. This represents an outstanding basis for future cooperation between MAX IV and HZB.”
The development of these substance libraries was funded by the German Federal Ministry of Education and Research (BMBF) and took place within the framework of the collaborative research project Frag4Lead.
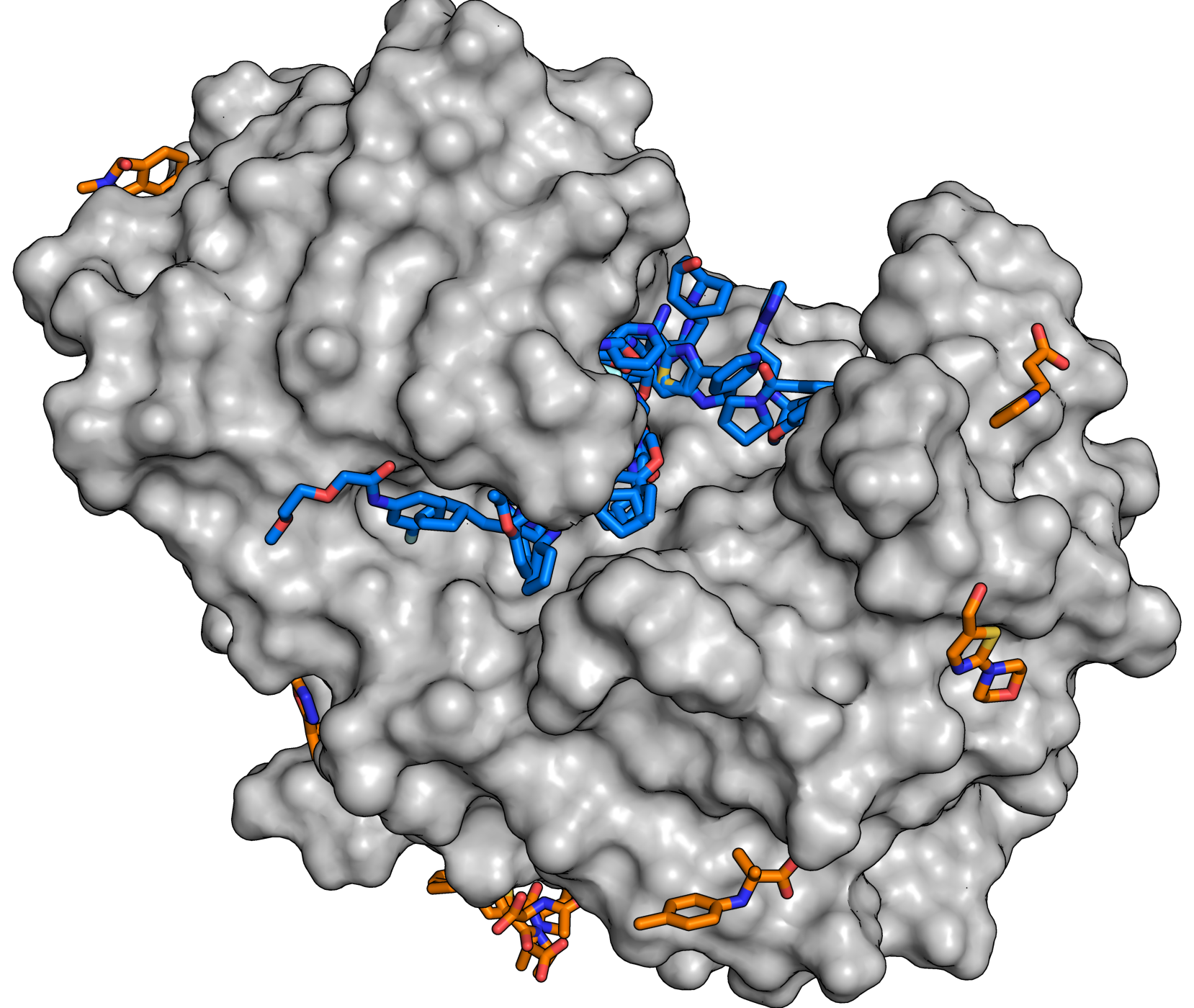
Title illustration: For the study, the enzyme endothiapepsin (grey) was brought into contact with molecules from the fragment library. The analyses now show that a large number of substances (blue and orange molecules) dock to the enzyme and each substance that is found is a potential starting point for the development of larger molecules. © J. Wollenhaupt/HZB