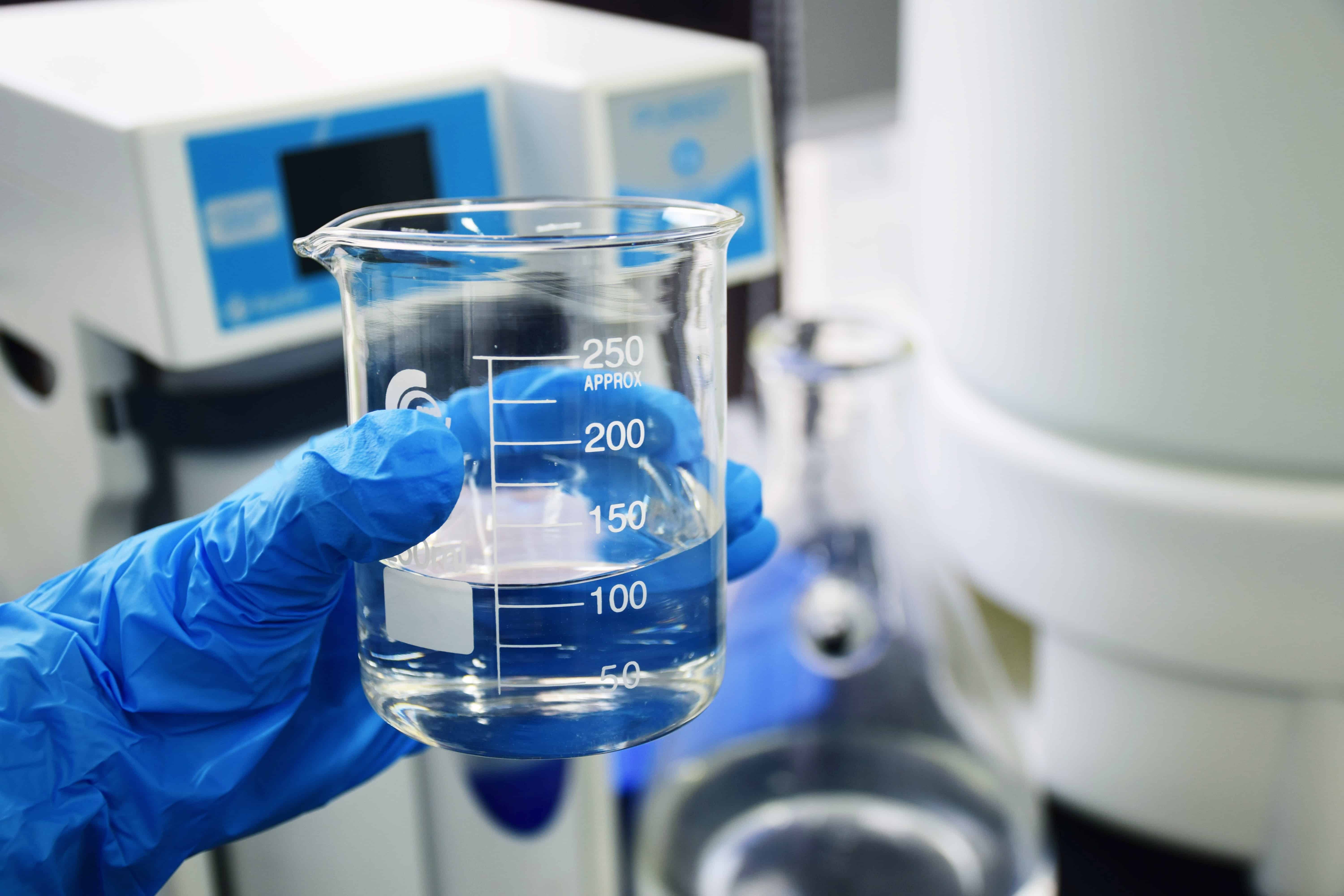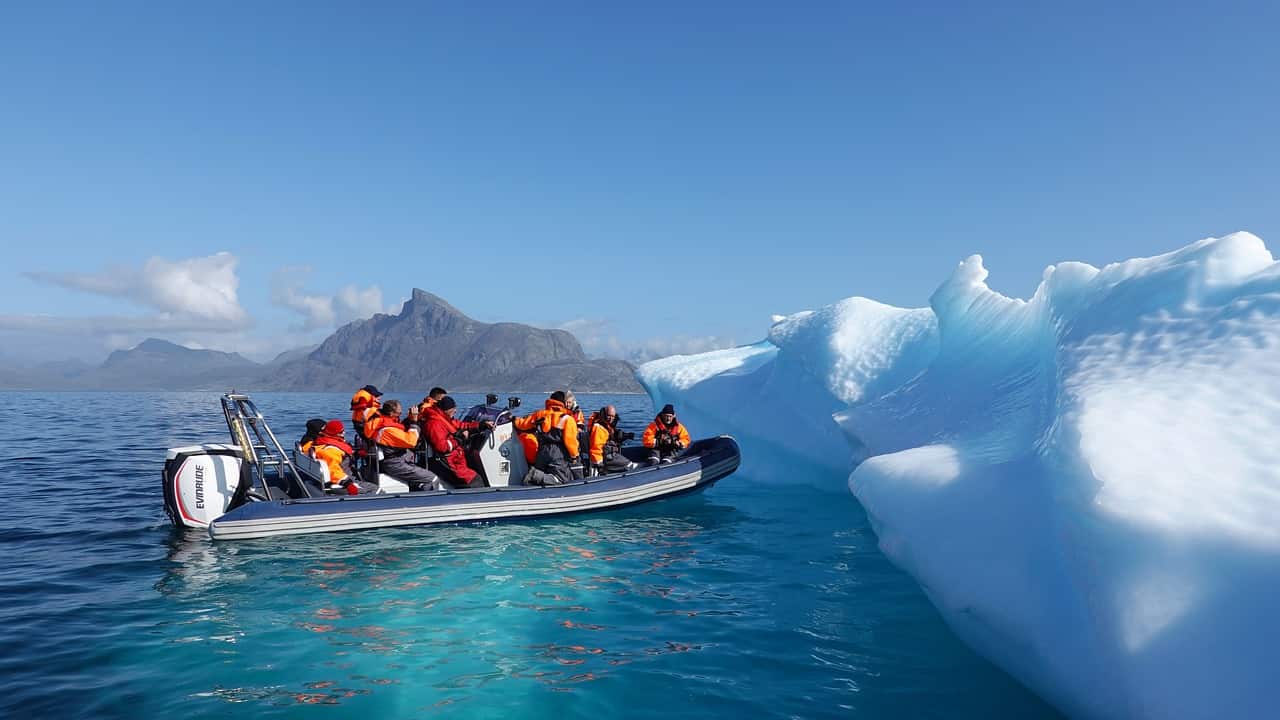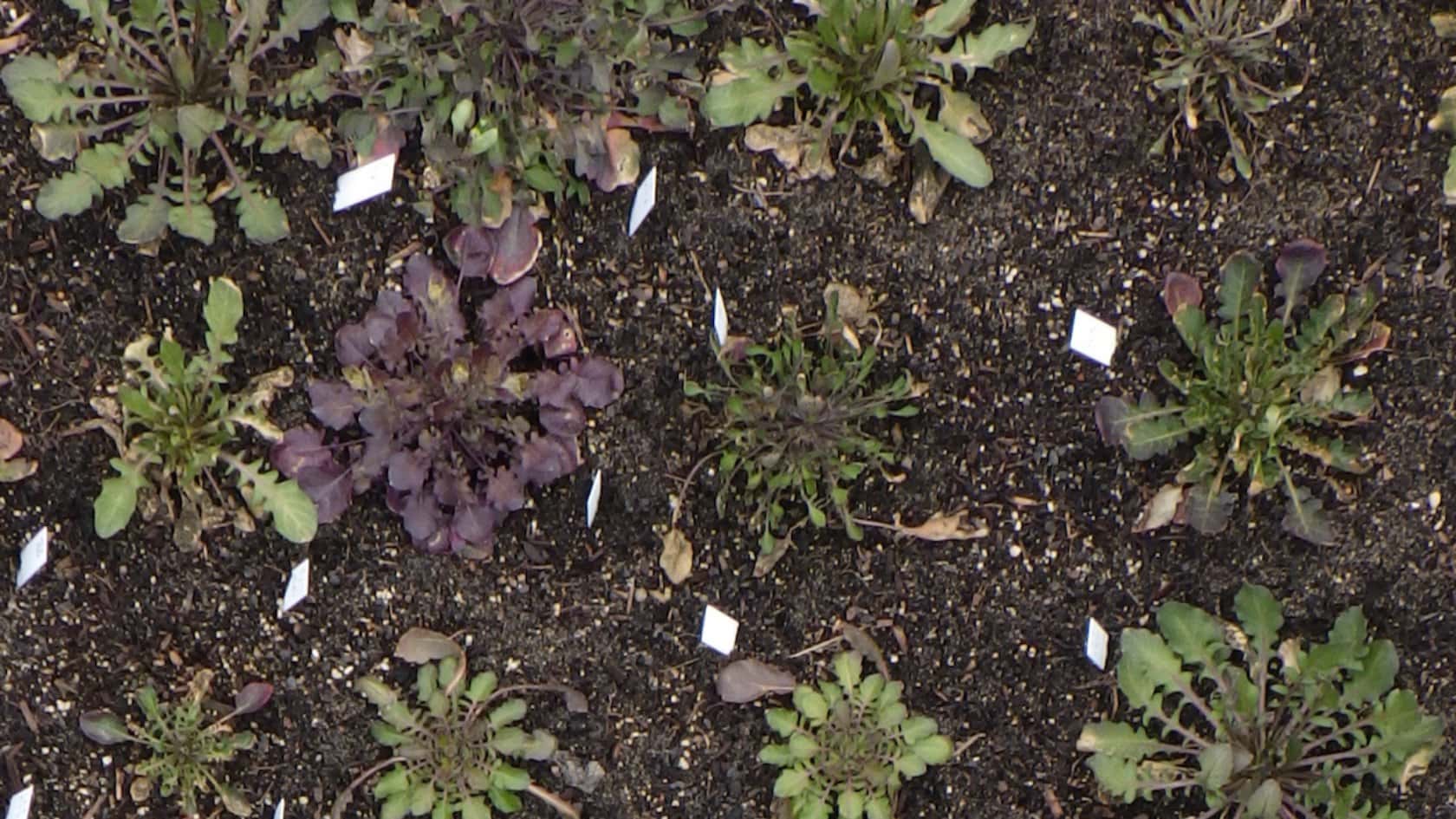
GenoRobotics (Lausanne), a project backed by École Polytechnique Fédérale de Lausanne (EPFL), is developing and testing a portable device for taking DNA measurements. They are developing it so they can take on-the-spot DNA samples of plant species and map biodiversity, even as climate change causes it to shift. The problem with current methods is that DNA often needs to be stored and sent to a specialized lab in order to be characterized – which can be difficult to do when taking samples in remote places. The samples can also be tainted along the way, meaning one would have to go back to get another sample.
This device would allow scientists to extract and sequence DNA while in the field, allowing them to create more accurate maps of changes to region specific flora.
GenoRobotics
Backed by EPFL’s MAKE program, the GenoRobotics project is headed by two EPFL graduates, Nicolas Adam and Jonathan Selz. The two came up with the idea while on a research expedition in Madagascar to characterize lemur environments. The samples had to be sent back to the United States for analysis but – due to administrative red-rape – the samples took two years before they could be analysed in a lab. The two realized they needed to find a way to sequence and analyse DNA to avoid these lengthy delays.
To speed up the extraction process, the technology uses a hydrogel developed by EPFL students. The hydrogel was originally designed to administer vaccines and contains tiny needles. It is a small, square piece of gel that – once pressed on a species – can use the needles to extract the DNA.
Our tests show that strands obtained this way are pure enough and concentrated enough to perform DNA barcoding,” says Adam in , in an article by EPFL. “To prevent the samples from becoming contaminated as they’re extracted, we’re developing a new mechanism that works like a stapler.”
Genome database
The two are also looking to create a data base that can be used and accessed in areas where internet connections are unreliable. Sequencing a genome can require dozens of gigabytes of memory and the two plan to use DNA barcoding – short segments of DNA – where a single gene can allow for characterization. Once analysed, the idea is to upload them to the database where it can interface with existing, publicly available databases iBOL and GenBank.







